Google Scholar (Link)
Pubmed (Link)
BioRxiv pre-prints (Link)
2026
Structural modeling reveals phage proteins that manipulate bacterial immune signaling.
Tal N, Hadary R, Chang RB, Osterman I, Jacobson R, Yirmiya E, Bechon N, Hochhauser D, López Rivera M, Madhala B, Garb J, Goldsmith M, Wein T, Kranzusch PJ, Amitai G, Sorek R.
Science. 2026; 391(6789), eaea1761. PMID 41785351.
Nucleotide signals coordinate activation and inhibition of bacterial immunity.
Yamaguchi S, Fernandez SG, Wassarman DR, Lüders M, Schwede F, Kranzusch PJ.
Nature. 2026; Advanced online publication. PMID 41708866.
Giant DNA viruses encode a hallmark translation initiation complex of eukaryotic life.
Fels JM, Hill AB, Han R, Bisio H, Abergel C, Kranzusch PJ, Lee ASY.
Cell. 2026; 189(5), 1423–1433. PMID 41709453.
A prophage-encoded abortive infection protein preserves host and prophage spread.
Sargen MR, Antine SP, Grabe GJ, Antonellis G, Ragucci AE, Li Y, Kranzusch PJ, Helaine S.
Nature. 2026; Advanced online publication. PMID 41606329.
2025
Mechanisms of herpesvirus helicase–primase inhibition and replication fork complex assembly.
Yu Z, Sathyanarayana P, Liu C, Tan JMJ, Yang P, Das B, Hu S, Fan X, Ji C, Weller SK, Shekhar M, Coen DM,
Kranzusch PJ, Loparo JJ, Abraham JA.
Cell. 2025; Advanced online publication. PMID 41468884.
Deazaguanylation is a nucleobase-protein conjugation required for type IV CBASS immunity.
Wassarman DR, Pfaff P, Paulo JA, Gygi SP, Shokat KM, Kranzusch PJ.
Science. 2025; 389(6767), 1347–1352. PMID 40997174.
A widespread family of viral sponge proteins reveals specific inhibition of nucleotide signals in anti-phage defense.
Chang RB, Toyoda HC, Hobbs SJ, Richmond-Buccola D, Wein T, Burger N, Chouchani ET, Sorek R, Kranzusch PJ.
Molecular Cell. 2025; 85(16), 3151–3165. PMID 40845805.
A DNA-gated molecular guard controls bacterial Hailong anti-phage defence.
Tan JMJ, Melamed S, Cofsky JC, Syangtan D, Hobbs SJ, Marmol JD, Jost M, Kruse AC, Sorek R, Kranzusch PJ.
Nature. 2025; 643(8072), 794–800. PMID 40306316.
TIR domains produce histidine-ADPR as an immune signal in bacteria.
Sabonis D*, Avraham C*, Chang RB*, Lu A, Herbst E, Silanskas A, Vilutis D, Leavitt A, Yirmiya E, Toyoda HC, Ruksenaite R, Zaremba M, Osterman I, Amitai G, Kranzusch PJ#, Sorek R#, Tamulaitiene G#.
Nature. 2025; 642(8067), 467–473. PMID 40307559. (*co-first, #co-corresponding)
CARD domains mediate anti-phage defence in bacterial gasdermin systems.
Wein T, Millman A, Lange K, Yirmiya E, Hadary R, Garb J, Melamed S, Amitai G, Dym O, Steinruecke F, Hill AB, Kranzusch PJ#, Sorek R#.
Nature. 2025; 639(8005), 727–734. PMID 39880956. (#co-corresponding)
Cell-free assays reveal that the HIV-1 capsid protects reverse transcripts from cGAS immune sensing.
Scott TM, Arnold LM, Powers JA, McCann DA, Rowe AB, Christensen DE, Pereira MJ, Zhou W, Torrez RM, Iwasa JH, Kranzusch PJ, Sundquist WI, Johnson JS.
PLoS Pathog. 2025; 21(1):e1012206. PMID 39874383.
Structure-guided discovery of viral proteins that inhibit host immunity.
Yirmiya E, Hobbs SJ, Leavitt A, Osterman I, Avraham C, Hochhauser D, Madhala B, Skovorodka M, Tan JMJ, Toyoda HC, Chebotar I, Itkin M, Malitsky S, Amitai G, Kranzusch PJ, Sorek R.
Cell. 2025; 188(6), 1681–1692. PMID 39855193.
2024
Animal and bacterial viruses share conserved mechanisms of immune evasion.
Hobbs SJ, Nomburg J, Doudna JA, Kranzusch PJ.
Cell. 2024; 187(20), 5530–5539. PMID 39197447.
Nucleotide immune signaling in CBASS, Pycsar, Thoeris, and CRISPR antiphage defense.
Hobbs SJ, Kranzusch PJ.
Annual Review in Microbiology. 2024; 78(1), 255–276. PMID 39083849.
A large-scale type I CBASS antiphage screen identifies the phage prohead protease as a key determinant of immune activation and evasion.
Richmond-Buccola D, Hobbs SJ, Garcia JM, Toyoda H, Gao J, Shao S, Lee ASY, Kranzusch PJ.
Cell Host & Microbe. 2024; 32(7), 1074–1088. PMID 38917809.
The SARM1 TIR domain produces glycocyclic ADPR molecules as minor products.
Garb J*, Amitai G*, Lu A, Ofir G, Brandis A, Mehlman T, Kranzusch PJ, Sorek R.
PLOS One. 2024; 19(4), e0302251. PMID 38635746.
Structure and assembly of a bacterial gasdermin pore.
Johnson AG#, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ#.
Nature. 2024; 628(8008), 657–663. PMID 38509367. (#co-corresponding)
Tracing the evolutionary origins of antiviral immunity.
Eaglesham JB, Kranzusch PJ.
PLOS Biology. 2024; 22(2), e3002481. PMID 38319913.
Structural basis of Gabija anti-phage defence and viral immune evasion.
Antine SP, Johnson AG, Mooney SE, Leavitt A, Mayer ML, Yirmiya E, Amitai G, Sorek R, Kranzusch PJ.
Nature. 2024; 625(7994), 360–365. PMID 37992757.
Phages overcome bacterial immunity via diverse anti-defence proteins.
Yirmiya E*, Leavitt A*, Lu A, Ragucci AE, Avraham C, Osterman I, Garb J, Antine SP, Mooney SE, Hobbs SJ, Kranzusch PJ, Amitai G#, Sorek R#.
Nature. 2024; 625(7994), 352–359. PMID 37992756. (*co-first, #co-corresponding)
2023
Bacterial SEAL domains undergo autoproteolysis and function in regulated intramembrane proteolysis.
Brogan AP, Habib C, Hobbs SJ, Kranzusch PJ, Rudner DZ.
PNAS. 2023; 120(40), e2310862120. PMID 37756332.
RNase L-activating 2′–5′ oligoadenylates bind ABCF1, ABCF3, and Decr-1.
Govande AA*, Babnis AW*, Urban C, Habjan M, Hartmann R, Kranzusch PJ#, Pichlmair A#.
Journal of General Virology. 2023; 104(9), 001890. PMID 37676257. (*co-first, #co-corresponding)
A virus-induced cyclic dinucleotide 2′3′-c-di-GMP mediates STING-dependent antiviral immunity in Drosophila.
Cai H*#, Li L*, Slavik KM*, Huang J, Yin T, Ai X, Hédelin L, Haas G, Xiang Z, Yang Y, Li X, Chen Y, Wei Z, Deng H, Chen D, Jiao R, Martins N, Meignin C, Kranzusch PJ#, Imler JL.
Immunity. 2023; 56, 1991–2005. PMID 37659413. (*co-first, #co-corresponding)
Structural homology screens reveal host-derived poxvirus protein families impacting inflammasome activity.
Boys IN, Johnson AG, Quinlan MR, Kranzusch PJ, Elde NC.
Cell Reports. 2023; 42(8), 112878. PMID 37494187.
Editorial Overview: Evolution of antiviral defense.
Kranzusch PJ.
Current Opinion in Microbiology. 2023; 75, 102352. PMID 37406561.
CBASS to cGAS-STING: the origins and mechanisms of nucleotide second messenger immune signaling.
Slavik KM, Kranzusch PJ.
Annual Review in Virology. 2023; 10(1), 423–453. PMID 37380187.
cGLRs are a diverse family of pattern recognition receptors in animal innate immunity.
Li Y, Slavik KM, Toyoda HC, Morehouse BR, de Oliveira Mann CC, Elek A, Levy S, Wang Z, Mears KS, Liu J, Kashin D, Guo X, Mass T, Sebé-Pedrós A, Schwede F, Kranzusch PJ.
Cell. 2023; 186(15), 3261–3276. PMID 37379839.
Viral sponges sequester nucleotide signals to inactivate immunity.
Richmond-Buccola D, Kranzusch PJ.
Trends in Microbiology. 2023; 31(6), 552–553. PMID 37100632.
Cryo-EM structure of the RADAR supramolecular anti-phage defense complex.
Duncan-Lowey B*, Tal N*, Johnson AG, Rawson S, Mayer ML, Doron S, Millman A, Melamed S, Fedorenko T, Kacen A, Brandis A, Mehlman T, Amitai G, Sorek R#, Kranzusch PJ#.
Cell. 2023; 186(5), 987–998. PMID 36764290. (*co-first, #co-corresponding)
Species-specific self-DNA detection mechanism by mammalian cyclic GMP-AMP synthases.
Mosallanejad K, Kennedy SN, Bahleda KM, Slavik KM, Zhou W, Govande AA, Hancks DC, Kranzusch PJ, Kagan JC..
Science Immunology. 2023; 8(79), eabp9765. PMID 36662885.
2022
A microbial transporter of the dietary antioxidant ergothioneine.
Dumitrescu DG, Gordon EM, Kovalyova Y, Seminara AB, Duncan-Lowey B, Forster ER, Zhou W, Booth CJ, Shen A, Kranzusch PJ, Hatzios S.
Cell. 2022; 185(24), 4526–4540. PMID 36347253.
What bacterial cell death teaches us about life.
Johnson AG, Kranzusch PJ.
PLOS Pathogens. 2022; 18(10), e1010879. PMID 36301823.
Sequential action of a tRNA base editor in conversion of cytidine to pseudouridine.
Kimura S#, Srisuknimit V, McCarty K, Dedon PC, Kranzusch PJ, Waldor MK#.
Nature Communications. 2022; 13(1), 5994. PMID 36220828. (#co-corresponding)
Viruses inhibit TIR gcADPR signalling to overcome bacterial defence.
Leavitt A*, Yirmiya E*, Amitai G*, Lu A*, Garb J, Herbst E, Morehouse BR, Hobbs SJ, Antine SP, Sun ZYJ, Kranzusch PJ#, Sorek R#.
Nature. 2022; 611(7935), 326–331. PMID 36174646. (*co-first, #co-corresponding)
Structural basis of human TREX1 DNA degradation and autoimmune disease.
Zhou W#, Richmond-Buccola D, Wang Q, Kranzusch PJ#.
Nature Communications. 2022; 13(1), 4277. PMID 35879334. (#co-corresponding)
Cryo-EM structure of an active bacterial TIR-STING filament complex.
Morehouse BR, Yip MCJ, Keszei AFA, McNamara-Bordewick NK, Shao S#, Kranzusch PJ#.
Nature. 2022; 608(7924), 803–807. PMID 35859168. (#co-corresponding)
Phage anti-CBASS and anti-Pycsar nucleases subvert bacterial immunity.
Hobbs SJ, Wein T, Lu A, Morehouse BR, Schnabel J, Leavitt A, Yirmiya E, Sorek R, Kranzusch PJ.
Nature. 2022; 605(7910), 522–526. PMID 35395152.
CBASS phage defense and evolution of antiviral nucleotide signaling.
Duncan-Lowey B, Kranzusch PJ.
Current Opinion in Immunology. 2022; 74, 156–163. PMID 35123147.
Bacterial gasdermins reveal an ancient mechanism of cell death.
Johnson AG*, Wein T*, Mayer ML, Duncan-Lowey B, Yirmiya E, Oppenheimer-Shaanan Y, Amitai G, Sorek R#, Kranzusch PJ#.
Science. 2022; 375(6577), 221–225. PMID 35025633. (*co-first, #co-corresponding)
2021
Effector-mediated membrane disruption controls cell death in CBASS antiphage defense.
Duncan-Lowey B, McNamara-Bordewick NK, Tal N, Sorek R, Kranzusch PJ.
Molecular Cell. 2021; 81(24), 5039–5051. PMID 34784509.
Cyclic CMP and cyclic UMP mediate bacterial immunity against phages.
Tal N*, Morehouse BR*, Millman A, Stokar-Avihail A, Avraham C, Fedorenko T, Yirmiya E, Herbst E, Brandis A, Mehlman T, Oppenheimer-Shaanan Y, Keszei AFA, Shao S, Amitai G, Kranzusch PJ#, Sorek R#.
Cell. 2021; 184(23), 5728–5739. PMID 34644530. (*co-first, #co-corresponding)
cGAS-like receptors sense RNA and control 3′2′-cGAMP signalling in Drosophila.
Slavik KM, Morehouse BR, Ragucci AE, Zhou W, Ai X, Chen Y, Li L, Wei Z, Bähre H, König M, Seifert R, Lee ASY, Cai H, Imler JL, Kranzusch PJ.
Nature. 2021; 597(7874), 109–113. PMID 34261127.
Molecular basis of CD-NTase nucleotide selection in anti-phage defense.
Govande AA, Duncan-Lowey B, Eaglesham JB, Whiteley AT, Kranzusch PJ.
Cell Reports. 2021; 35, 109206. PMID 34077735.
cGAS phase separation inhibits TREX1-mediated DNA degradation and enhances cytosolic DNA sensing.
Zhou W, Mohr L, Maciejowski J, Kranzusch PJ.
Molecular Cell. 2021; 81(4), 739–755. PMID 33606975.
2020
Structures of diverse poxin cGAMP nucleases reveal a widespread role for cGAS-STING evasion in host-pathogen conflict.
Eaglesham JB, McCarty KL, Kranzusch PJ.
eLife. 2020; 9:e59753. PMID 33191912.
STING cyclic dinucleotide sensing originated in bacteria.
Morehouse BR, Govande AA, Millman A, Kezei AFA, Lowey B, Ofir G, Shao S, Sorek R, Kranzusch PJ.
Nature. 2020; 586(7829), 429–433. PMID 32877915.
A novel STING1 variant causes a recessive form of STING-associated vasculopathy with onset in infancy (SAVI).
Lin B, Berard R, Al Rasheed A, Aladba B, Kranzusch PJ, Henderlight M, Grom A, Kahle D, Torreggiani S, Aue AG, Mitchell J, de Jesus AA, Schulert GS, Goldbach-Mansky R.
J Allergy Clin Immunol. 2020; 146(5), 1204–1208. PMID 32673614.
CBASS immunity uses CARF-related effectors to sense 3′–5′- and 2′–5′-linked cyclic oligonucleotide signals and protect bacteria from phage infection.
Lowey B, Whiteley AT, Keszei AFA, Morehouse BR, Mathews IT, Antine SP, Cabrera VJ, Kashin D, Niemann P, Jain M, Schwede F, Mekalanos JJ, Shao S, Lee ASY, Kranzusch PJ.
Cell. 2020; 182(1), 38–49. PMID 32544385.
Conserved strategies for pathogen evasion of cGAS-STING immunity.
Eaglesham JB, Kranzusch PJ.
Current Opinion in Immunology. 2020; 66, 27–34. PMID 32339908.
Structure and mechanism of a cyclic trinucleotide-activated bacterial endonuclease mediating bacteriophage immunity.
Lau RK*, Ye Q*, Birkholz EA, Berg KR, Patel L, Matthews IT, Watrous JD, Ego K, Whiteley AT, Lowey B, Mekalanos JJ, Kranzusch PJ, Jain M, Pogliano J, Corbett KD.
Molecular Cell. 2020; 77(4), 723–733. PMID 31932164. (*co-first)
2019
cGAS and CD-NTase enzymes: structure, mechanism, and evolution.
Kranzusch PJ.
Current Opinion in Structural Biology. 2019; 59, 178–187. PMID 31593902.
Analysis of human cGAS activity and structure.
Zhou W, Whiteley AT, Kranzusch PJ.
Methods in Enzymology. 2019; 625, 13–40. PMID 31455523.
Modular architecture of the STING C-terminal tail allows interferon and NF-κB signaling adaptation.
de Oliveira Mann CC, Orzalli MH, King DS, Kagan JC, Lee ASY#, Kranzusch PJ#.
Cell Reports. 2019; 27, 1165–1175. PMID 31018131. (#co-corresponding)
Structure and mechanism of a Hypr GGDEF enzyme that activates cGAMP signaling to control extracellular metal respiration.
Hallberg ZF*, Chan CH*, Wright TA, Kranzusch PJ, Doxzen KW, Park JJ, Bond DR#, Hammond MC#.
eLife. 2019; 8:e43959. PMID 30964001. (*co-first, #co-corresponding)
Phosphoinositide interactions position cGAS at the plasma membrane to ensure efficient distinction between self- and viral DNA.
Barnett KC, Coronas-Serna JM, Zhou W, Ernandes MJ, Cao A, Kranzusch PJ, Kagan JC.
Cell. 2019; 176(6), 1432–1446. PMID 30827685.
Bacterial cGAS-like enzymes synthesize diverse nucleotide signals.
Whiteley AT, Eaglesham JB, de Oliveira Mann CC, Morehouse BR, Lowey B, Nieminen EA, Danilchanka O, King DS, Lee ASY,
Mekalanos JJ#, Kranzusch PJ#.
Nature. 2019; 567(7747), 194–199. PMID 30787435. (#co-corresponding)
Viral and metazoan poxins are cGAMP-specific nucleases that restrict cGAS-STING signalling.
Eaglesham JB, Pan Y, Kupper TS, Kranzusch PJ.
Nature. 2019; 566(7743), 259–263. PMID 30728498.
Recurrent loss-of-function mutations reveal costs to OAS1 antiviral activity in primates.
Carey CM, Govande AA, Cooper JM, Hartley MK, Kranzusch PJ, Elde NC.
Cell Host & Microbe. 2019; 25(2), 336–343. PMID 30713099.
2018
Structure of the human cGAS–DNA complex reveals enhanced control of immune surveillance.
Zhou W*, Whiteley AT*, de Oliveira Mann CC, Morehouse BR, Nowak RP, Fischer ES, Gray NS, Mekalanos JJ, Kranzusch PJ.
Cell. 2018; 174(2), 300–311. PMID 30007416. (*co-first)
2017
cGAS Conducts Micronuclei DNA Surveillance.
de Oliveira Mann CC, Kranzusch PJ.
Trends in Cell Biology. 2017; 27(10), 697698. PMID 28882413.
Dynamic control of X-chromosome conformation and repression by a histone H4K20 demethylase.
Brejc K*, Bian Q*, Uzawa S, Wheeler BS, Anderson EC, King DS, Kranzusch PJ, Preston CG, Meyer BJ.
Cell. 2017; 171(1), 85–102. PMID 28867287. (*co-first)
A broad-spectrum inhibitor of CRISPR-Cas9.
Harrington LB*, Doxzen KW*, Ma E, Liu J, Knott GJ, Edraki A, Garcia B, Amrani N, Chen JS, Cofsky JC, Kranzusch PJ, Sontheimer EJ, Davidson AR, Maxwell KL, Doudna JA.
Cell. 2017; 170(6), 1224–1233. PMID 28844692. (*co-first)
-------- Start of lab at Harvard Medical School in 2016 --------
2016
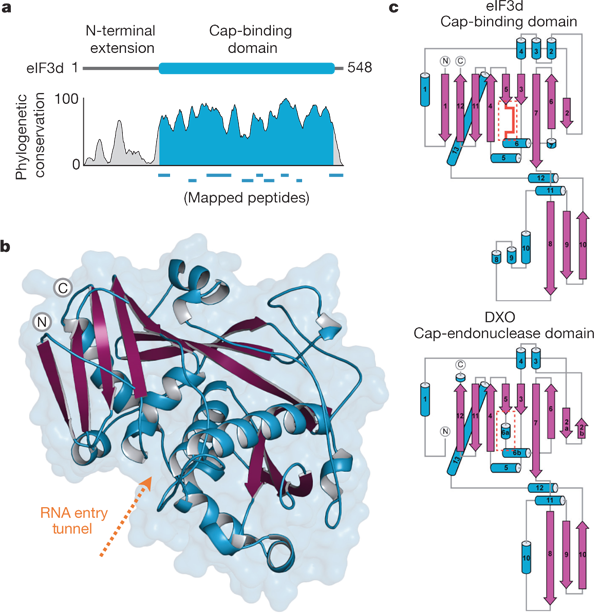
eIF3d is an mRNA cap-binding protein required for specialized translation initiation.
Lee AS, Kranzusch PJ, Doudna JA, Cate JD.
Nature. 2016; 536(7614), 96–9. PMID 27462815.
A bacterial Argonaute with noncanonical guide RNA specificity.
Kaya E*, Doxzen KW*, Knoll KR, Wilson RC, Strutt SC, Kranzusch PJ, Doudna JA.
PNAS. 2016; 113(15), 4057–62. PMID 27035975. (*co-first)
2015
Foreign DNA capture during CRISPR-Cas adaptive immunity.
Nuñez JK*, Harrington LB*, Kranzusch PJ, Engelman AN, Doudna JA.
Nature. 2015; 527(7579), 535–8. PMID 26503043. (*co-first)
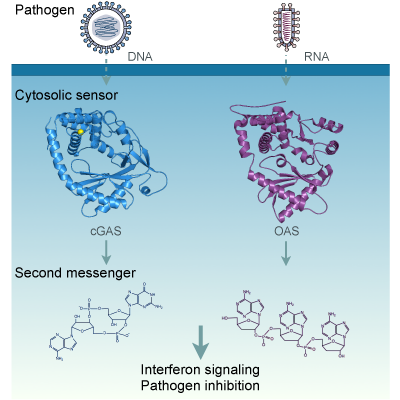
Ancient Origin of cGAS-STING Reveals Mechanism of Universal 2',3' cGAMP Signaling.
Kranzusch PJ*, Wilson SC*, Lee AS, Berger JM, Doudna JA#, Vance RE#.
Molecular Cell. 2015; 59(6), 891–903. PMID 26300263. (*co-first, #co-corresponding)
eIF3 targets cell-proliferation messenger RNAs for translational activation or repression.
Lee AS, Kranzusch PJ, Cate JH.
Nature. 2015; 522(7554), 111–4. PMID 25849773.
2014
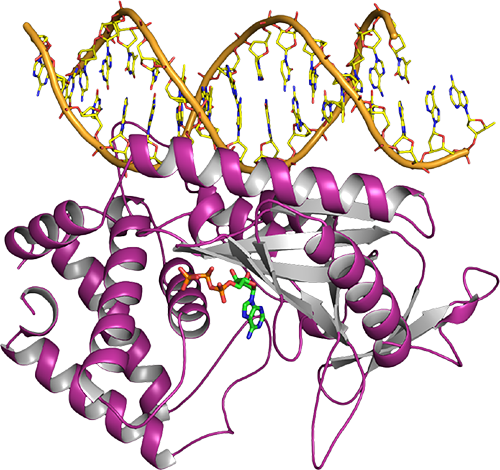
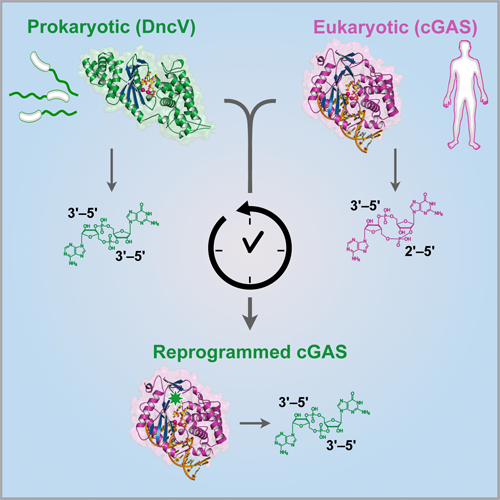
Structure-guided reprogramming of human cGAS dinucleotide linkage specificity.
Kranzusch PJ, Lee AS, Wilson SC, Solovykh MS, Vance RE, Berger JM#, Doudna JA#.
Cell. 2014; 158(5), 1011–21. PMID 25131990. (#co-corresponding)
Cas1-Cas2 complex formation mediates spacer acquisition during CRISPR-Cas adaptive immunity.
Nuñez JK, Kranzusch PJ, Noeske J, Wright AV, Davies CW, Doudna JA.
Nature Structural & Molecular Biology. 2014; 21(6), 528–34. PMID 24793649.
African origin of the malaria parasite Plasmodium vivax.
Liu W, Li Y, Shaw KS, Learn GH, Plenderleith LJ, Malenke JA, Sundararaman SA, Ramirez MA, Crystal PA, Smith AG, Bibollet-Ruche F, Ayouba A, Locatelli S, Esteban A, Mouacha F, Guichet E, Butel C, Ahuka-Mundeke S, Inogwabini BI, Ndjango JB, Speede S, Sanz CM, Morgan DB, Gonder MK, Kranzusch PJ, Walsh PD, Georgiev AV, Muller MN, Piel AK, Stewart FA, Wilson ML, Pusey AE, Cui L, Wang Z, Färnert A, Sutherland CJ, Nolder D, Hart JA, Hart TB, Bertolani P, Gillis A, LeBreton M, Tafon B, Kiyang J, Djoko CF, Schneider BS, Wolfe ND, Mpoudi-Ngole E, Delaporte E, Carter R, Culleton RL, Shaw GM, Rayner JC, Peeters M, Hahn BH, Sharp PM.
Nature Communications. 2014; 5, 3346. PMID 24557500.
2013
cGAS dimerization entangles DNA recognition.
Kranzusch PJ, Vance RE.
Immunity. 2013; 39(6), 992–4. PMID 24332024.
Structure of human cGAS reveals a conserved family of second-messenger enzymes in innate immunity.
Kranzusch PJ, Lee AS, Berger JM#, Doudna JA#.
Cell Reports. 2013; 3(5), 1362–8. PMID 23707061. (#co-corresponding)
The polymerase of negative-stranded RNA viruses.
Morin B, Kranzusch PJ, Rahmeh AA, Whelan SP.
Current Opinion in Virology. 2013; 3(2), 103–10. PMID 23602472.
2012
Architecture and regulation of negative-strand viral enzymatic machinery.
Kranzusch PJ, Whelan SP.
RNA Biology. 2012; 9(7), 941–8. PMID 22767259.
2011
Arenavirus Z protein controls viral RNA synthesis by locking a polymerase-promoter complex.
Kranzusch PJ, Whelan SP.
PNAS. 2011; 108(49), 19743–8. PMID 22106304.
Ebola virus entry requires the cholesterol transporter Niemann-Pick C1.
Carette JE*, Raaben M*, Wong AC*, Herbert AS, Obernosterer G, Mulherkar N, Kuehne AI, Kranzusch PJ, Griffin AM, Ruthel G, Dal Cin P, Dye JM#, Whelan SP#, Chandran K#, Brummelkamp TR#.
Nature. 2011; 477(7364), 340–3. PMID 21866103. (*co-first, #co-corresponding)
Ebolavirus delta-peptide immunoadhesins inhibit marburgvirus and ebolavirus cell entry.
Radoshitzky SR, Warfield KL, Chi X, Dong L, Kota K, Bradfute SB, Gearhart JD, Retterer C, Kranzusch PJ, Misasi JN, Hogenbirk MA, Wahl-Jensen V, Volchkov VE, Cunningham JM, Jahrling PB, Aman MJ, Bavari S, Farzan M, Kuhn JH.
Journal of Virology. 2011; 85(17), 8502–13. PMID 21697477.
2010
Molecular architecture of the vesicular stomatitis virus RNA polymerase.
Rahmeh AA, Schenk AD, Danek EI, Kranzusch PJ, Liang B, Walz T, Whelan SP.
PNAS. 2010; 107(46), 20075–80. PMID 21041632.
Assembly of a functional Machupo virus polymerase complex.
Kranzusch PJ, Schenk AD, Rahmeh AA, Radoshitzky SR, Bavari S, Walz T, Whelan SP.
PNAS. 2010; 107(46), 20069–74. PMID 20978208.
Origin of the human malaria parasite Plasmodium falciparum in gorillas.
Liu W, Li Y, Learn GH, Rudicell RS, Robertson JD, Keele BF, Ndjango JB, Sanz CM, Morgan DB, Locatelli S, Gonder MK, Kranzusch PJ, Walsh PD, Delaporte E, Mpoudi-Ngole E, Georgiev AV, Muller MN, Shaw GM, Peeters M, Sharp PM, Rayner JC, Hahn BH.
Nature. 2010; 467(7314), 420–5. PMID 20864995.
Infectious Lassa virus, but not filoviruses, is restricted by BST-2/tetherin.
Radoshitzky SR, Dong L, Chi X, Clester JC, Retterer C, Spurgers K, Kuhn JH, Sandwick S, Ruthel G, Kota K, Boltz D, Warren T, Kranzusch PJ, Whelan SP, Bavari S.
Journal of Virology. 2010; 84(20), 10569–80. PMID 20686043.
Molecular epidemiology of simian immunodeficiency virus infection in wild-living gorillas.
Neel C, Etienne L, Li Y, Takehisa J, Rudicell RS, Bass IN, Moudindo J, Mebenga A, Esteban A, Van Heuverswyn F, Liegeois F, Kranzusch PJ, Walsh PD, Sanz CM, Morgan DB, Ndjango JB, Plantier JC, Locatelli S, Gonder MK, Leendertz FH, Boesch C, Todd A, Delaporte E, Mpoudi-Ngole E, Hahn BH, Peeters M.
Journal of Virology. 2010; 84(3), 1464–76. PMID 19906908.
2009
Ribose 2'-O methylation of the vesicular stomatitis virus mRNA cap precedes and facilitates subsequent guanine-N-7 methylation by the large polymerase protein.
Rahmeh AA, Li J, Kranzusch PJ, Whelan SP.
Journal of Virology. 2009; 83(21), 11043–50. PMID 19710136.